Here is the application of the n-gram model to text genereation using the full text of the play "Hamlet" as training data:
text = ExampleData[{"Text", "Hamlet"}];
genTexts = {#,
NGramMarkovChainText[text, #,
StringSplit[text][[1020 ;; 1020 + # - 1]], 200,
WordSeparators -> {" ", "\n"}]} & /@ Range[2, 5]
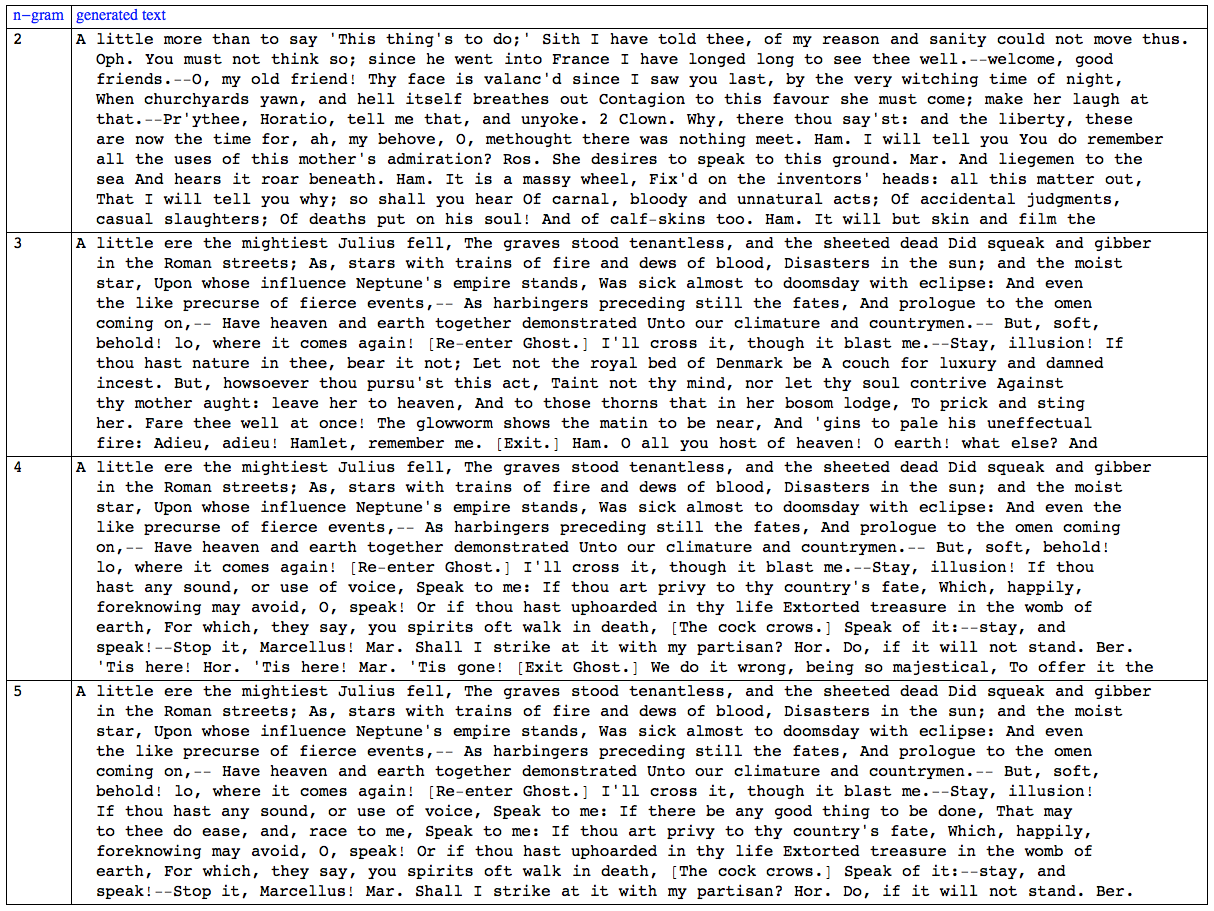
It can be seen in the table that the 5-gram generated text makes more sense than the 2-gram one. All 4 randomly generated texts start from the same place in the play.
(A more detailed discussion is given in my "
Markov chains n-gram model implementation" blog post.)