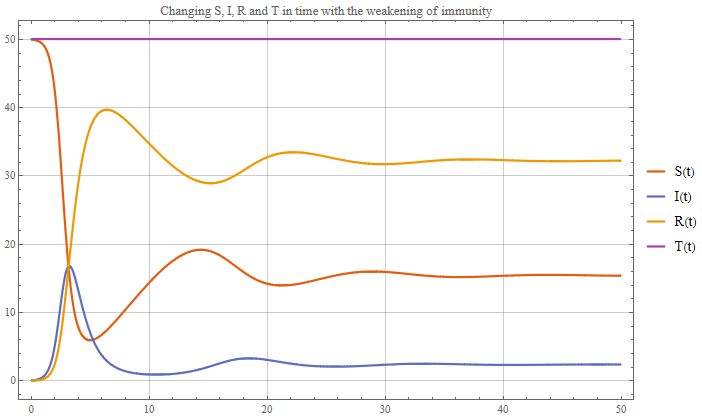
Hi Arnoud! I was asked for help by a student with a similar problem. There was a similar model, but it differed in that the loss of immunity over time was added. Those. those who fell ill once could get sick again. Actually below is the code and graphics.
Params1 = {T0 -> 50.05, a -> 0.07, b -> 1.08, c -> 0.08};
Conditions1 = {T[0] == T0, II[0] == T0*0.1/100, R[0] == 0};
SIRm = {
S'[t] == -a*S[t]*II[t] + c*R[t],
II'[t] == a*S[t]*II[t] - b*II[t],
R'[t] == b*II[t] - c*R[t],
T[t] == S[t] + II[t] + R[t]
};
{Smsol, IImsol, Rmsol, Tmsol} =
First[{S, II, R, T} /.
NDSolve[Join[SIRm, Conditions1] /. Params1, {S, II, R, T}, {t, 0,
50}]];
Plot[{Smsol[t], IImsol[t], Rmsol[t], Tmsol[t]}, {t, 0, 50},
PlotTheme -> "Scientific", ImageSize -> Large,
PlotLegends -> {"S(t)", "I(t)", "R(t)", "T(t)"},
GridLines -> Automatic, PlotStyle -> Thick,
PlotLabel ->
"Changing S, I, R and T in time with the weakening of immunity"]