Last challenge for this week: From Paul Wellin's "Essentials of Programming in Mathematica"
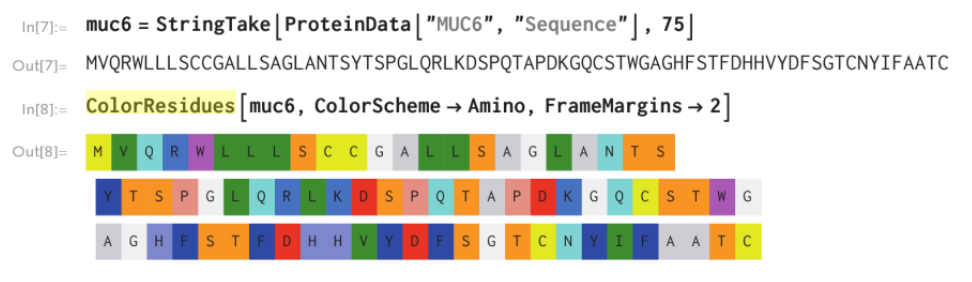
Create a function ColorResidues[seq] that takes an amino acid sequence seq and returns a visualization like that below where the amino acids are colored according to a scheme such as Amino, where more polar residues are brighter and more nonpolar residues are colored darker. In addition to a color scheme option add options to control the frame around each residue.
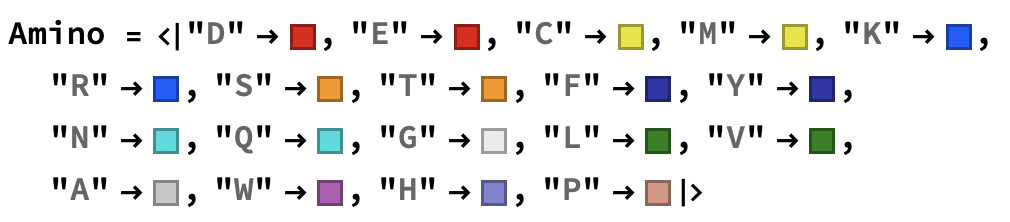
Note: Changing Amino to an Association. I think that will work better. Also for reference, I got the colors from http://life.nthu.edu.tw/~fmhsu/rasframe/SHAPELY.HTM
Amino = <|"D" -> RGBColor["#E60A0A"],
"E" -> RGBColor["#E60A0A"],
"C" -> RGBColor["#E6E600"],
"M" -> RGBColor["#E6E600"],
"K" -> RGBColor["#145AFF"],
"R" -> RGBColor["#145AFF"],
"S" -> RGBColor["#FA9600"],
"T" -> RGBColor["#FA9600"],
"F" -> RGBColor["#3232AA"],
"Y" -> RGBColor["#3232AA"],
"N" -> RGBColor["#00DCDC"],
"Q" -> RGBColor["#00DCDC"],
"G" -> RGBColor["#EBEBEB"],
"I" -> RGBColor["#0F820F"],
"L" -> RGBColor["#0F820F"],
"V" -> RGBColor["#0F820F"],
"A" -> RGBColor["#C8C8C8"],
"W" -> RGBColor["#B45AB4"],
"H" -> RGBColor["#8282D2"],
"P" -> RGBColor["#DC9682"]|>
muc6 = StringTake[ProteinData["MUC6", "Sequence"], 75]